| Source |
Annotation |
Motif |
Evidence |
| HT-SELEX | Yin2017 | 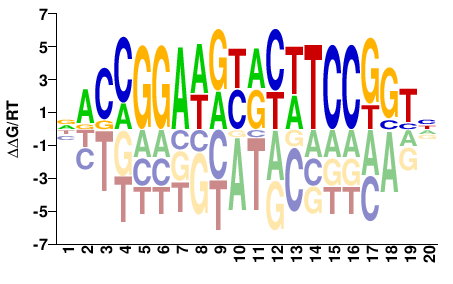 | Direct |
| Methyl-HT-SELEX | Yin2017 | 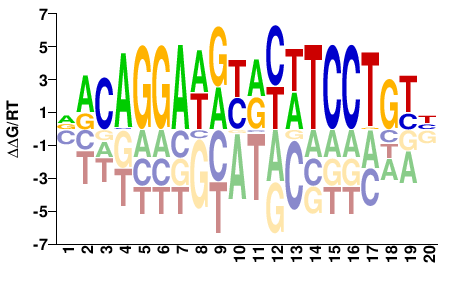 | Direct |
| Transfac | Transfac | License required | Direct |
| Transfac | Transfac | License required | Direct |
| Misc | HocoMoco |  | Direct |
| PBM | Wei10 | 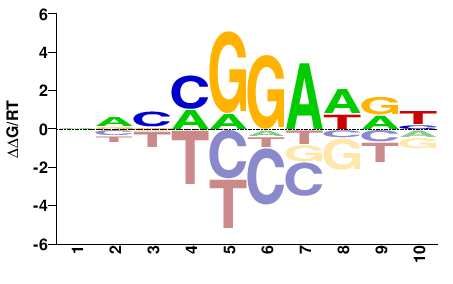 | Inferred - Ets1 (96% AA Identity, Mus musculus) |
| Transfac | Transfac | License required | Inferred - ETS1B_CHICK (96% AA Identity, Gallus gallus) |
| HT-SELEX | Nitta2015 |  | Inferred - pnt (93% AA Identity, Drosophila melanogaster) |
| ChIP-seq | ENCODE | 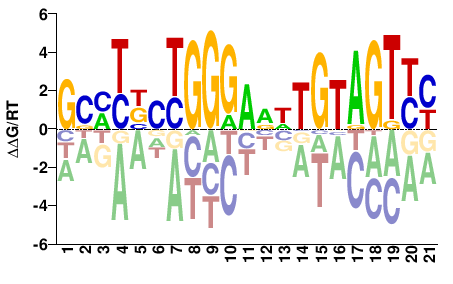 | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| ChIP-seq | ENCODE |  | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| SELEX | Hallikas2006 |  | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| SELEX | JASPAR |  | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Misc | HocoMoco |  | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| Misc | HOMER |  | Inferred - ETS1 (90% AA Identity, Homo sapiens) |
| SMiLE-seq | Isakova2017 |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| PBM | Badis09 |  | Inferred - Gabpa (67% AA Identity, Mus musculus) |
| PBM | Wei10 |  | Inferred - Gabpa (67% AA Identity, Mus musculus) |
| ChIP-seq | ENCODE |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| ChIP-seq | ENCODE |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| ChIP-seq | ENCODE |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| ChIP-seq | ENCODE |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| ChIP-seq | ENCODE |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| ChIP-seq | JASPAR |  | Inferred - Gabpa (67% AA Identity, Mus musculus) |
| Misc | HocoMoco |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| Misc | HOMER |  | Inferred - GABPA (67% AA Identity, Homo sapiens) |
| HT-SELEX | Jolma2010 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| HT-SELEX | Jolma2010 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| HT-SELEX | Nitta2015 |  | Inferred - Ets65A (65% AA Identity, Drosophila melanogaster) |
| HT-SELEX | Yin2017 |  | Inferred - ETV1 (65% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ETV1 (65% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV1 (65% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV1 (65% AA Identity, Homo sapiens) |
| PBM | Wei10 |  | Inferred - Etv1 (65% AA Identity, Mus musculus) |
| PBM | Wei10 |  | Inferred - Etv1 (65% AA Identity, Mus musculus) |
| SELEX | JASPAR |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ERG (65% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ERG (65% AA Identity, Homo sapiens) |
| Misc | HocoMoco |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| Misc | HOMER |  | Inferred - ERG (65% AA Identity, Homo sapiens) |
| Misc | HOMER |  | Inferred - ETV1 (65% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| PBM | Wei10 |  | Inferred - Elk1 (64% AA Identity, Mus musculus) |
| PBM | Wei10 |  | Inferred - Erg (64% AA Identity, Mus musculus) |
| HT-SELEX | Nitta2015 |  | Inferred - Ets21C (64% AA Identity, Drosophila melanogaster) |
| PBM | Zoo_01 |  | Inferred - Ets21C (64% AA Identity, Drosophila melanogaster) |
| HT-SELEX | Yin2017 |  | Inferred - ETV5 (64% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV5 (64% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV5 (64% AA Identity, Homo sapiens) |
| PBM | Wei10 |  | Inferred - Etv5 (64% AA Identity, Mus musculus) |
| HT-SELEX | Yin2017 |  | Inferred - FEV (64% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - FEV (64% AA Identity, Homo sapiens) |
| SMiLE-seq | Isakova2017 |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| PBM | Wei10 |  | Inferred - Fli1 (64% AA Identity, Mus musculus) |
| SELEX | JASPAR |  | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| SELEX | JASPAR |  | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| Misc | HocoMoco |  | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| Misc | HOMER | 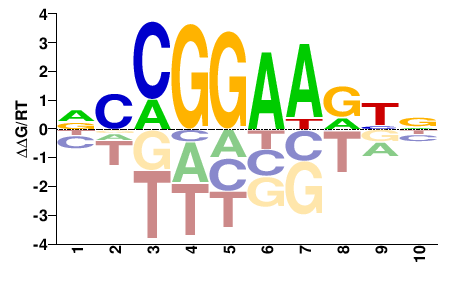 | Inferred - ELK1 (64% AA Identity, Homo sapiens) |
| Misc | HocoMoco | 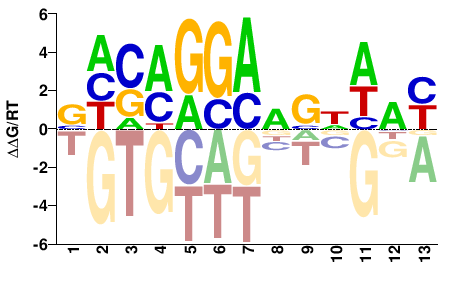 | Inferred - ETV5 (64% AA Identity, Homo sapiens) |
| Misc | HocoMoco |  | Inferred - FEV (64% AA Identity, Homo sapiens) |
| Misc | HocoMoco | 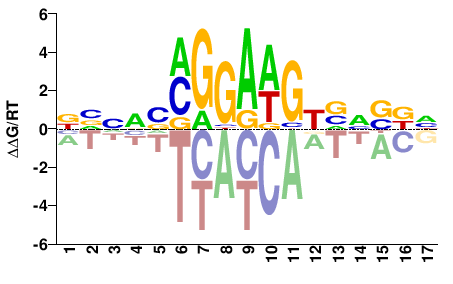 | Inferred - FLI1 (64% AA Identity, Homo sapiens) |
| Misc | HOMER | 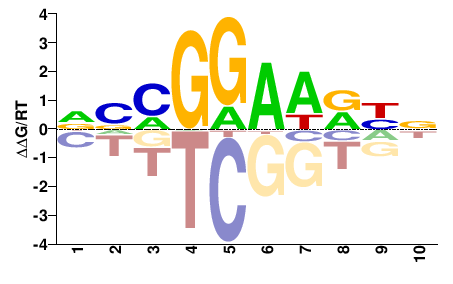 | Inferred - Fli1 (64% AA Identity, Mus musculus) |
| HT-SELEX | Nitta2015 |  | Inferred - Ets96B (63% AA Identity, Drosophila melanogaster) |
| PBM | Zoo_01 |  | Inferred - elk4 (62% AA Identity, Tetraodon nigroviridis) |
| HT-SELEX | Yin2017 |  | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| PBM | Wei10 |  | Inferred - Etv4 (62% AA Identity, Mus musculus) |
| Transfac | Transfac | License required | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| Misc | HocoMoco |  | Inferred - ETV4 (62% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ETV2 (60% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV2 (60% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ETV2 (60% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ELK3 (59% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ELK3 (59% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ELK3 (59% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ELK3 (59% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ELK3 (59% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ELK3 (59% AA Identity, Homo sapiens) |
| PBM | Wei10 |  | Inferred - Elk3 (59% AA Identity, Mus musculus) |
| Transfac | Transfac | License required | Inferred - ELK3 (59% AA Identity, Homo sapiens) |
| Misc | HocoMoco |  | Inferred - ELK3 (59% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ELK4 (58% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ELK4 (58% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ELK4 (58% AA Identity, Homo sapiens) |
| PBM | Wei10 |  | Inferred - Elk4 (58% AA Identity, Mus musculus) |
| PBM | Wei10 | 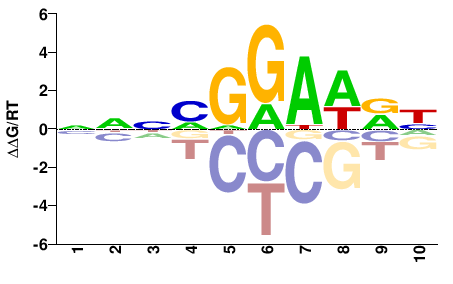 | Inferred - XP_911724.4 (58% AA Identity, Mus musculus) |
| ChIP-seq | ENCODE |  | Inferred - ELK4 (58% AA Identity, Homo sapiens) |
| ChIP-seq | ENCODE |  | Inferred - ELK4 (58% AA Identity, Homo sapiens) |
| ChIP-seq | JASPAR |  | Inferred - ELK4 (58% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ELK4 (58% AA Identity, Homo sapiens) |
| Transfac | Transfac | License required | Inferred - ELK4 (58% AA Identity, Homo sapiens) |
| Misc | HocoMoco |  | Inferred - ELK4 (58% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ETV3 (56% AA Identity, Homo sapiens) |
| HT-SELEX | Yin2017 |  | Inferred - ETV3 (56% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 |  | Inferred - ETV3 (56% AA Identity, Homo sapiens) |
| Methyl-HT-SELEX | Yin2017 | 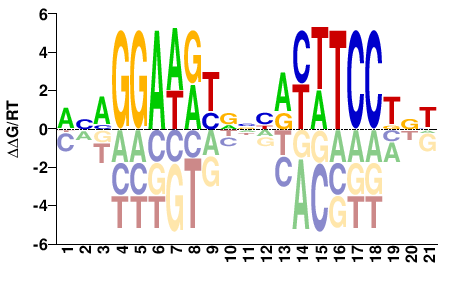 | Inferred - ETV3 (56% AA Identity, Homo sapiens) |
| PBM | Wei10 | 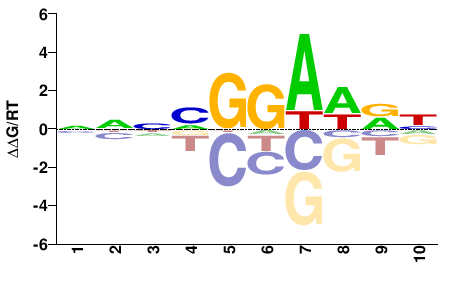 | Inferred - Etv3 (56% AA Identity, Mus musculus) |